Amplicon Barcoding
The main purpose of this protocol is to add Illumina adapters via a 2-step PCR. This is mainly used for sequencing of targeted regions of a genome and metagenomics. It is a general method for generating libraries from PCR products.
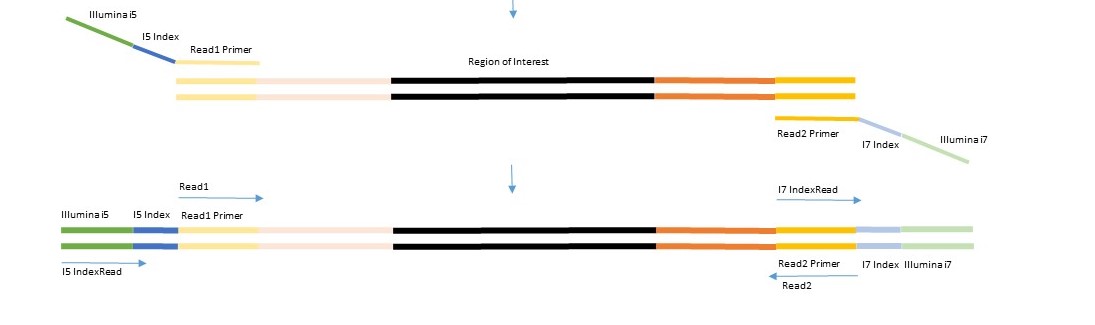
The 1st PCR step is to amplify the region of interest (ROI) with ROI specific primers that contain part of the Illumina adapter sequence. This PCR is done by the user.

The 2nd PCR step is to add the second part of the Illumina adapter which contain the indices. The second primer will bind to Illumina adapter sequence of the first PCR. This will result in barcoded ready-to-sequence libraries.
We made an excel sheet to help you order the 1st PCR primers properly. You can download it here:
IlluminaAmpliconPrimerColorCoded.xlsx
This image explains how to use it
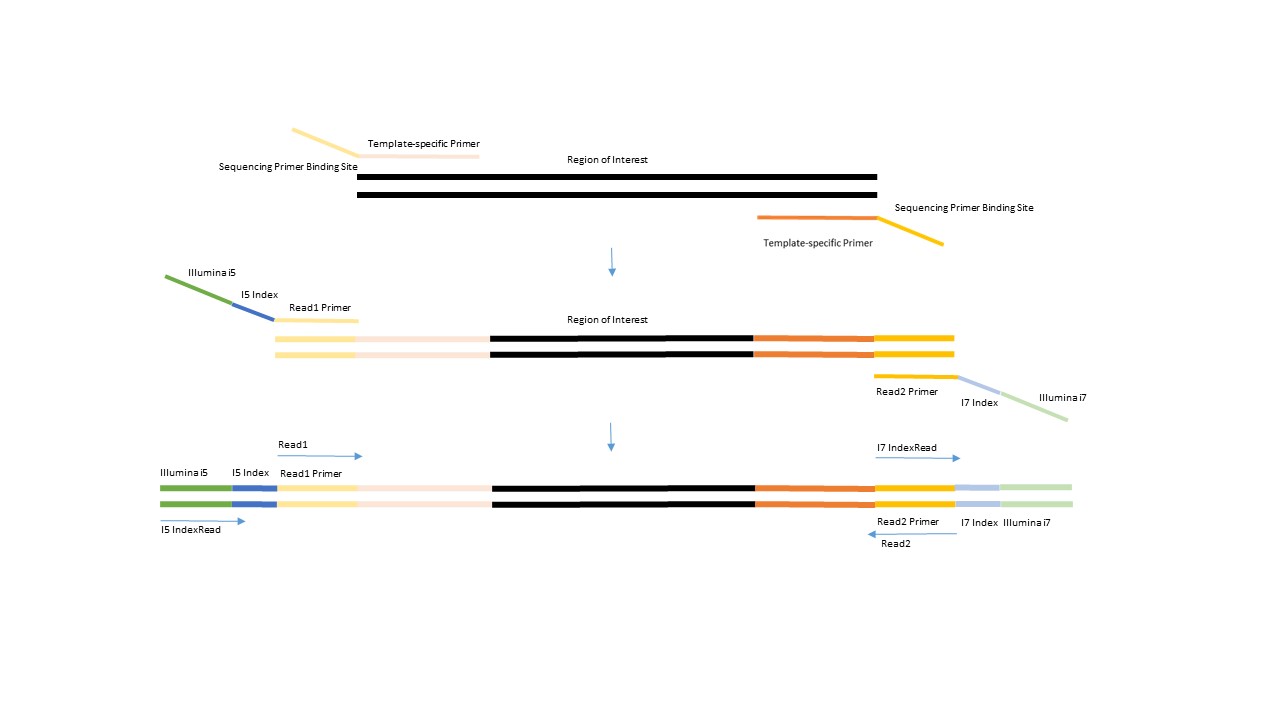
This is a summary of the Amplicon generation process:
| 
|
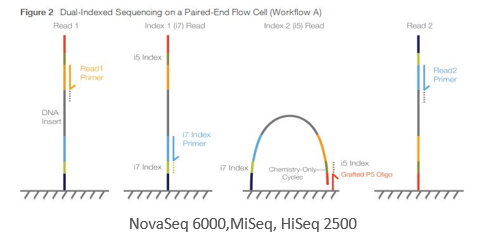
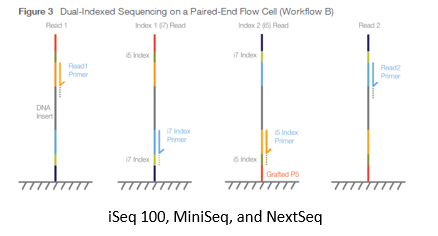
Figure2,3 are from Document # 15057455 v04 of Illumina Inc
Back to Short Read/Illumina Page
Back to FGCZ Main Page